Leveraging Contrastive Learning and Domain Prompts for Efficient Radiology Report Generation
Abstract
Generating radiology report, especially for chest X-rays, remains a crucial yet time-consuming task in clinical practice. Although recent AI frameworks show promise, they face significant challenges including poor long-form generation, content hallucination, and the requirement of massive training datasets. To address these challenges, we propose a novel CLIP-based framework that incorporates a modified contrastive le\pt tuning mechanism that adapts BioBERT to radiology-specific terminology while preserving pre-trained knowledge. Furthermore, we implement an enhanced DenseNet121 architecture for improved feature extraction, particularly for rare pathological conditions. Our experimental evaluation on the IU X-ray dataset demonstrates state-of-the-art performance, achieving BLEU-1, ROUGE, and METEOR scores of 0.48, 0.37, and 0.22, respectively.
초록
흉부 X-ray 영상에 대한 방사선 판독문 생성은 임상 진료에서 매우 중요하지만 시간이 많이 소요되는 작업으로 남아있다. 최근 AI 프레임워크들이 유망한 결과를 보여주고 있지만, 긴 문장 생성의 한계, 내용 환각, 그리고 대규모 학습 데이터 요구와 같은 중요한 도전과제들이 여전히 존재한다. 이러한 문제들을 해결하기 위해, 본 연구는 변형된 대조 학습 방식을 통합하여 학습 데이터 요구사항을 줄일 수 있는 새로운 CLIP 기반 프레임워크를 제안한다. 제안한 프레임워크는 사전 학습된 지식을 유지하면서도 BioBERT를 방사선학 특화 용어에 적응시키는 소프트 프롬프트 튜닝 메커니즘을 특징으로 한다. 더불어, 희귀 병리 상태에 대한 특징 추출을 개선하기 위해 향상된 DenseNet121 아키텍처를 구현하였다. IU X-ray 데이터셋에 대한 실험 평가에서 제안한 방법은 BLEU-1, ROUGE, METEOR 점수에서 각각 0.48, 0.37, 0.22를 달성하며 최첨단 성능을 보여주었다.
Keywords:
deep learning, contrastive learning, soft prompt tuning, chest x-raysⅠ. Introduction
Medical imaging technologies, such as chest X-rays, are widely used in various diagnostic and treatment scenarios. They enable physicians to accurately identify the causes of patients’ conditions, facilitating the development of effective treatment plans. However, generating accurate and comprehensive medical reports is a complex process that demands medical expertise and substantial diagnostic experience. This task consumes a considerable amount of time and effort from physicians, posing challenges in the face of increasing diagnostic workloads. The World Health Organization(WHO) anticipates a global shortage of 18 million healthcare professionals by 2030, significantly affecting the number of radiology specialists[1]. Consequently, there is an urgent need to develop efficient and accurate methods for generating medical reports to enhance the productivity and quality of healthcare services. This growth has sparked significant research interest in computer-aided diagnostic and treatment technologies in recent years.
One promising solution that has received considerable attention is automatic radiology report generation, which aims to reduce the heavy workload faced by radiologists. Most modern methods utilize an encoder-decoder framework, wherein medical images (e.g., chest X-rays) are first transformed into latent feature representations using Convolutional Neural Networks(CNNs)[2][3]. These representations are then decoded into natural language sentences to produce radiology reports, employing Recurrent Neural Networks(RNNs)[4] or fully attentive architectures such as Transformers[5].
Despite current achievements, two primary challenges persist: (1) extracting comprehensive and clinically relevant information from the medical images and (2) ensuring accurate Cross-Modal Alignments(CMA), which involve linking generated textual content to corresponding regions within the images.
On the other hand, contrastive learning offers a compelling approach to address some of these challenges, particularly in the context of learning image-text pairs, such as X-ray images and their corresponding textual reports[6]. One of its key advantages is its ability to learn representations from large volumes of unlabeled data. It is particularly beneficial in medical domains where annotated datasets are often limited due to the need for expert labeling. Contrastive learning leverages the similarities and differences across image-text pairs to enhance model performance using abundant unlabeled datasets. However, recent contrastive learning approaches are often data-intensive and may struggle with small-scale datasets, a common constraint in the medical domain[7].
In this work, we propose a novel automatic radiology report generation approach that combines CLIP-based image representations with soft prompt tuning on BioBERT. This combination addresses the challenges associated with limited data availability and the demands of large-scale models. By employing soft prompt tuning, we effectively reduce the risk of overfitting on smaller datasets while utilizing the extensive medical knowledge encoded in the BioBERT to ensure accurate report generation. Simultaneously, we enhance the DenseNet121 architecture with an attention and pooling mechanism tailored to improve the extraction and interpretation of features from chest X-ray images. This dual focus on robust image representation and precise cross-modal alignment ensures the generation of radiology reports that are both accurate and contextually relevant. By integrating these techniques, we aim to provide a scalable, efficient, and reliable solution to support healthcare professionals in addressing the increasing diagnostic demands.
The main contributions of this work are summarized as follows:
1. We present an effective approach for leveraging the potential of large-scale contrastive learning models in the medical domain, specifically addressing the challenges of limited annotated data.
2. Our method involves a soft prompt tuning method, which facilitates efficient knowledge transfer from large-scale BioBERT model while reducing the risk of overfitting.
3. Additionally, we design a novel attention and pooling mechanism within the DenseNet121 architecture, enhancing its capability to learn and interpret anatomical structures for precise feature extraction in radiology report generation.
The remainder of this paper is organized as follows: Section II reviews related work in radiology report generation and contrastive learning methods. Section III details methodology, including the soft prompt tuning approach and the modified DenseNet121 architecture. Section IV presents experimental results and performance evaluations, followed by a comprehensive discussion and ablation studies in Section V. Finally, Section VI concludes the paper with insights and future directions.
Ⅱ. Related Work
Radiology report generation has gained significant attention in recent years, with many methods leveraging the encoder-decoder architecture originally developed for image captioning tasks. However, generating radiology reports poses unique challenges compared to image captioning, as medical reports are typically longer and identification of clinical abnormalities in chest X-rays is more complex due to inherent data biases in training datasets. To address these challenges, researchers have proposed a variety of innovative contributions to traditional methods.
For instance, the Text-Image Embedding Network(TieNet) was proposed[8] to extract distinctive image and text representations using a multilevel attention mechanism integrated into an end-to-end trainable CNN-RNN architecture. TieNet first classifies chest X-rays by combining image features with text embeddings extracted from corresponding reports and then uses attention mechanisms to generate detailed reports. Building on this, another approach employs a CNN encoder coupled with multi-stage RNNs as decoders[9] demonstrating improved efficiency in translating medical images into reports compared to traditional RNN models. Medical reports often contain heterogeneous information, including paragraphs, tags, and keywords, presenting additional challenges. To address this, a multi-task framework[10] was introduced to perform tag and paragraph generation simultaneously. LSTM-based models are used to produce long and diverse medical report paragraphs. However, while RNN and LSTM models have been widely adopted for medical report generation, they suffer from inefficiencies in generating longer, coherent texts[11].
To overcome these limitations, transformer-based architectures with powerful attention mechanisms have become increasingly popular as decoders in radiology report generation tasks. For instance, a hierarchical transformer was proposed[12], integrating a CNN encoder to identify regions of interest using a bottom-up attention module, followed by a transformer decoder to produce coherent report paragraphs. To address the challenge of aligning key abnormalities with specific regions in chest X-rays, a Cross-modal Memory Network(CMN)[13] was introduced, enhancing the efficiency of transformer-based frameworks by incorporating shared memory, facilitating interaction and alignment between different modalities. Similarly, a memory-driven transformer[14] employed relational memory to capture crucial textual information during the generation process, integrating it into the decoder through memory-driven conditional layer normalization. Data bias presents another significant challenge in medical report generation, where normal visual regions often dominate over abnormal ones in X-ray images. To address this, the Align Transformer[15] was introduced, featuring the Align Hierarchical Attention(AHA) module, predicting disease tags from input images and hierarchically aligning these visual regions with disease tags.
While transformer-based models have demonstrated impressive accuracy in radiology tasks, the available Chest X-ray(CXR) datasets are often insufficient for training such computationally heavy models[20].
Recently, contrastive learning has emerged as a strong alternative to traditional encoder-decoder models. By learning the alignment between visual and textual data in an unsupervised manner, contrastive learning captures the inherent relationship between medical images and reports. ConVIRT[16], a notable technique in this domain, employs contrastive learning to maximize similarities between true image-text pairs while minimizing similarities for randomly generated negative pairs. This framework applies beyond report generation to other vision-language tasks, such as classification and image-to-text retrieval. Another innovative method, CXR-IRGen[17], utilizes Variational Autoencoders(VAEs) trained on contrastive embeddings derived from the MIMIC-CXR[18] dataset. While this approach uses encoders pre-trained on natural images, transferring them to medical report generation tasks can be challenging due to differences between natural and medical images, potentially limiting generalization in scenarios with limited data.
Ⅲ. Methodology
3.1 Overview
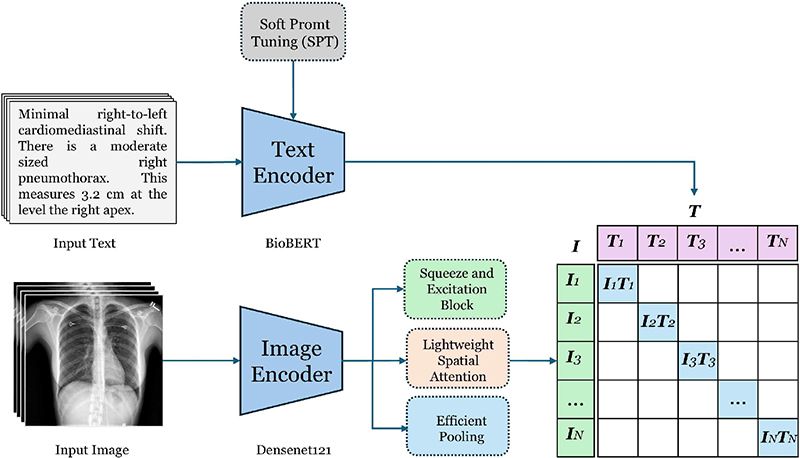
This study introduces a novel architecture for learning correlation between medical image-text pairs, building upon the fundamental principles of Contrastive Language-Image Pretraining(CLIP)[19], while introducing key modifications tailored for medical imaging applications. Our framework employs a dual-encoder architecture to analyze chest X-rays alongside their corresponding radiology reports, diverging from CLIP's original design through the strategic use of categorical cross-entropy loss instead of InfoNCE loss. This modification is particularly significant in the medical domain, where precise classification of pathological conditions requires more explicit supervision than general image-text matching.
While InfoNCE loss excels at learning broad image-text similarities through contrastive learning, categorical cross-entropy offers more detailed supervision by treating each image-text pair as a distinct class. This approach allows the model to capture subtle differences in medical conditions that might be overlooked in a purely contrastive approach.
The architecture comprises two primary components: a text encoder based on BioBERT, enhanced with prompt tuning capabilities, and a vision encoder based on DenseNet121 with advanced feature extraction mechanisms. These components are integrated into a cohesive multi-modal framework designed to learn joint representations of chest X-rays and their corresponding textual descriptions, as illustrated in Figure 1. This design choice enables our model to retain the powerful representation learning capabilities of CLIP while incorporating the precision and specificity required for medical imaging tasks.
3.2 Text encoder architecture
Our text encoder leverages a pre-trained BioBERT model enhanced with Soft Prompt Tuning(SPT) to effectively process clinical text data. BioBERT is chosen as the backbone due to its proven effectiveness in biomedical text processing, having been pre-trained on a vast corpus of PubMed abstracts and clinical notes. However, clinical text processing presents unique challenges, including domain-specific terminology, complex medical relationships, and limited labeled data, which require additional architectural solutions. The key innovation in our approach is the integration of SPT, which introduces learnable prompt embeddings to adapt the pre-trained BioBERT model to the medical domain while maintaining its original parameters unchanged. This approach offers several advantages:
(1) It significantly reduces the number of trainable parameters, making the model more efficient and less prone to overfitting
(2) It preserves the pre-trained knowledge while allowing task-specific adaptations
(3) It enables better few-shot learning capabilities, crucial for medical domains where labeled data is limited.
The text processing pipeline consists of several stages that progressively transform raw clinical text into rich, contextualized representations. Given a clinical report x, first, it passes through a specialized tokenization process using BioBERT's pre-trained tokenizer
| (1) |
Here, L represents the sequence length and d is the embedding dimension (468 for BioBERT). This initial embedding matrix E captures the basic semantic properties of each token while maintaining the sequential structure of the input text. Further with a set of P learnable prompt embeddings the token embeddings are enhanced, which are prepended to the token embeddings as given:
| (2) |
These prompt embeddings are initialized using a truncated normal distribution with a carefully chosen standard deviation of 0.02, by finding it empirically to provide stable training dynamics. Unlike traditional prompt tuning approaches, our prompts incorporate a dynamic scaling mechanism that adjusts the prompt influence based on the input sequence characteristics:
| (3) |
where α(x) is an attention-based scaling function that computes context-dependent importance weights for each prompt token. The enhanced embeddings are then processed through the frozen BioBERT model to obtain contextualized representations:
| (4) |
where H represents the final hidden states that capture local and global contextual information. The frozen nature of BioBERT during this process ensures that the pre-trained medical knowledge is preserved while allowing the prompt embeddings to guide the interpretation of the input text. To obtain a fixed-size representation of the entire clinical text, we employ a sophisticated pooling strategy that combines both global and local information:
| (5) |
This pooled representation captures the overall semantic content of the clinical text while maintaining sensitivity to important local details through the attention-weighted averaging process. The pooled representation undergoes a series of K projection layers that progressively refine the feature space. Let’s initial embeddings is represented as H0=HGlobal, then
| (6) |
where k∈{1, ..., k} and Wk∈R(d×d), Bk∈Rd are learnable parameters, and GELU is the Gaussian Error Linear Unit activation function. The residual connections (+Hk-1) ensure smooth gradient flow and prevent information loss during the projection process. Each projection layer is followed by layer normalization and dropout (rate = 0.1) to enhance training stability and prevent overfitting:
| (7) |
This multi-layer projection architecture allows the model to learn increasingly abstract representations while maintaining the medical domain-specific features captured by the prompt-tuned BioBERT encoder.
3.3 Vision encoder
This section describes the proposed vision encoder architecture for processing x-ray images. This architecture introduces several key innovations in feature extraction and attention mechanisms, building upon the established DenseNet121 architecture while incorporating novel approaches to handle the unique challenges of medical image analysis.
Given an input medical image , the network produces feature maps , preserving both local anatomical details and broader structural patterns. The dense connectivity pattern inherent in DenseNet121 ensures efficient feature reuse and gradient flow throughout the network, particularly important for the fine-grained analysis required in medical imaging.
Further, we implement parallel feature extraction pathways. These pathways process the initial features through depth-wise convolutions at different scales:
| (8) |
The features from these parallel pathways are then integrated through point-wise convolutions and concatenation:
| (9) |
where PW represents 1×1 convolution projecting features to d dimensions, and [;] denotes channel-wise concatenation.
A key innovation in our architecture is the dual-attention mechanism that separately processes channel and spatial relationships.
The channel attention mechanism implements a modified squeeze-and-excitation approach:
| (10) |
where and are learnable parameters (using reduction ratio r=4), δ and σ represent ReLU and sigmoid activations respectively, and GAP denotes global average pooling. This mechanism dynamically recalibrates channel-wise feature responses.
The spatial attention mechanism computes position-specific importance weights:
| (11) |
where Conv(7*7) represents a convolutional layer with 7×7 kernel size, producing a spatial attention map . This enables the network to focus on anatomically relevant regions.
Finally, we implement an adaptive pooling mechanism that learns to combine global average and max pooling operations:
| (12) |
where w∈[0,1] is a learned weight determining the relative contribution of each pooling operation, allowing the model to adapt its feature aggregation strategy based on image-specific characteristics. This adaptive combination allows us to leverage both the noise-reduction benefits of global average pooling and the discriminative feature preservation of max pooling, dynamically balancing these properties based on the input image characteristics.
3.4 Contrastive learning
The goal of contrastive learning is to learn aligned representations of text and images. Given a batch of N text-image pairs, we compute the pairwise cosine similarity between all text and image embeddings:
| (13) |
The cosine similarities are normalized using the softmax function to produce probabilities:
| (14) |
where τ is a temperature parameter that scales the logits. The loss function is the average categorical cross-entropy loss for both directions (text-to-image and image-to-text):
| (15) |
Here, Pi,i represents the probability of the correct text-image pair. Minimizing this loss ensures that embeddings of matched pairs are similar, while embeddings of mismatched pairs are dissimilar.
Ⅳ. Results
4.1 Datasets
In this study, the Indian University Chest X-rays(CXR) dataset is utilized which is widely recognized for radiology research, to train, validate, and test models. The dataset is available publicly and comprises 3,955 deidentified radiology reports and impressions, each paired with frontal and lateral CXR images, resulting in a total of 7,470 images. For this work, the dataset is divided into training, validation, and testing subsets in a 7:2:1 ratio. All model evaluations are conducted exclusively on the testing set to ensure unbiased performance assessment.
4.2 Comparison with state of the art models
The performance of our proposed framework is summarized in Table 1, which presents results based on the pre-split test data. The model supports retrieval tasks in both text-to-image and image-to-text formats.
To ensure a fair comparison with state-of-the-art models, we adopted an image-text retrieval approach. In this process, we first generated embeddings for all test images using the pre-trained vision encoder. Similarly, embeddings were created for all test reports as well as reports from the training data. Cosine similarity analysis was then performed between the test image embeddings and the report embeddings. Based on these similarity scores, the most relevant reports from the training set were retrieved for evaluation.
4.3 Qualitative results
We performed a qualitative analysis of the reports retrieved by our proposed framework for both normal and abnormal chest X-rays, as illustrated in Table 2. The retrieved results highlight the framework's ability to effectively differentiate normal cases from abnormal ones. Additionally, these qualitative results showcase the model's capacity to identify and accurately describe abnormalities, emphasizing its potential to capture critical clinical details in radiology reports.
Ⅴ. Discussion & Ablation
The development of the proposed framework involved a series of extensive experiments aimed at refining its core components. Two key innovations played a pivotal role in achieving the framework's success: soft prompt tuning and the modified DenseNet architecture. Soft prompt tuning addressed significant challenges in utilizing BioBERT for this task. Using BioBERT with frozen weights resulted in a lack of convergence, while fine-tuning all the weights led to overfitting on the small-scale dataset. By introducing soft prompt tuning, the framework reduced the number of trainable parameters, preserving the pre-trained knowledge of BioBERT while allowing task-specific adaptations. This approach enabled efficient learning and better generalization, particularly in scenarios with limited labeled data, striking a balance between efficiency and performance.
The modified DenseNet architecture introduced critical enhancements to improve the encoder's ability to analyze chest X-ray images effectively. The attention mechanism and pooling strategy were instrumental in capturing both global anatomical features and local details, enabling the model to focus on the most relevant areas of the images. This capability significantly enhanced the model's ability to identify key abnormalities in chest X-rays, addressing one of the primary challenges in radiology report generation.
5.1 Performance evaluation of modified densenet
To evaluate the effectiveness of the proposed DenseNet modifications, a comparative analysis was conducted against the baseline DenseNet model. The results demonstrated significant improvements across all performance metrics, with BLEU-4 score increased from 0.39 to 0.48, ROUGE score improved from 0.30 to 0.37, and similarly METEOR score from 0.16 to 0.22. These performance gains can be attributed to specific architectural advancements in the modified DenseNet.
The introduction of multi-scale feature extraction through parallel depthwise convolutions allowed the model to simultaneously capture fine-grained details and broader contextual patterns. This ensured a more comprehensive analysis of anatomical structures. Additionally, the dual attention mechanism combined channel-wise recalibration using squeeze-and-excitation with spatial attention. This allowed the model to prioritize informative feature channels and relevant spatial regions within the chest X-ray images. The pooling strategy further contributed to these improvements by introducing a learnable balance between Global Average Pooling(GAP) and Global Max Pooling(GMP). This ensured that both distributed and localized features were retained, which is crucial for generating accurate and detailed radiology reports.
Overall, these architectural modifications enhanced the model's ability to analyze complex anatomical features and relationships, establishing its superiority over the standard DenseNet architecture in radiology report generation tasks.
VI. Conclusion
To evaluate the effectiveness of the proposed DenseNet modifications, a comparative analysis was conducted against the baseline DenseNet model. The results demonstrated significant improvements across all performance metrics, with BLEU-4 score increased from 0.39 to 0.48, ROUGE score improved from 0.30 to 0.37, and similarly METEOR score from 0.16 to 0.22. These performance gains can be attributed to specific architectural advancements in the modified DenseNet. The introduction of multi-scale feature extraction through parallel depthwise convolutions allowed the model to simultaneously capture fine-grained details and broader contextual patterns. This ensured a more comprehensive analysis of anatomical structures.
Additionally, the dual attention mechanism combined channel-wise recalibration using squeeze-and-excitation with spatial attention. This allowed the model to prioritize informative feature channels and relevant spatial regions within the chest X-ray images. The pooling strategy further contributed to these improvements by introducing a learnable balance between GAP and GMP. This ensured that both distributed and localized features were retained, which is crucial for generating accurate and detailed radiology reports. Overall, these architectural modifications enhanced the model’s ability to analyze complex anatomical features and relationships, establishing its superiority over the standard DenseNet architecture in radiology report generation tasks.
Acknowledgments
This work was supported by Innovative Human Resource Development for Local Intellectualization program through the Institute of Information & Communications Technology Planning & Evaluation(IITP) grant funded by the Korea government(MSIT)(IITP-2025-RS-2022-00156287)
References
-
K. Konstantinidis, "The shortage of radiographers: A global crisis in healthcare", Journal of Medical Imaging and Radiation Sciences, Vol. 55, No. 4, pp. 101333, Jan. 2024.
[https://doi.org/10.1016/j.jmir.2023.10.001]

-
K. Simonyan and A. Zisserman, "Very deep convolutional networks for large-scale image recognition", arXiv:1409.1556, , Sep. 2014.
[https://doi.org/10.48550/arXiv.1409.1556]

-
K. He, X. Zhang, S. Ren, and J. Sun, "Deep residual learning for image recognition", 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, pp. 770-778, Jun. 2016.
[https://doi.org/10.1109/CVPR.2016.90]

-
S. Hochreiter and J. Schmidhuber, "Long short-term memory", Neural Computation, Vol. 9, No. 8, pp. 1735-1780, Nov. 1997.
[https://doi.org/10.1162/neco.1997.9.8.1735]

-
A. Vaswani, N. Shazeer, N. Parmar, J. Uszkoreit, L. Jones, A. N. Gomez, Ł. Kaiser, and I. Polosukhin, "Attention is all you need", Advances in Neural Information Processing Systems, Vol. 30, pp. 5998-6008, Dec. 2017.
[https://doi.org/10.48550/arXiv.1706.03762]

-
M. Endo, R. Wong, C. Suh, and P. Rajpurkar, "Retrieval-based chest X-ray report generation using a pre-trained contrastive language-image model", Machine Learning for Health, PMLR, pp. 209-220, Dec. 2021.
[https://doi.org/10.48550/arXiv.2109.12242]

-
A. Radford, et al., "Learning transferable visual models from natural language supervision", International Conference on Machine Learning, PMLR, pp. 8748-8763, Jul. 2021.
[https://doi.org/10.48550/arXiv.2103.00020]

-
X. Wang, Y. Peng, L. Lu, Z. Lu, M. Bagheri, and R. M. Summers, "TieNet: Text-image embedding network for common thorax disease classification and reporting in chest X-rays", IEEE/CVF Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, pp. 9049-9058, Jun. 2018.
[https://doi.org/10.1109/CVPR.2018.00943]

-
S. Singh, S. Karimi, K. Ho, and L. Torrey, "From chest X-rays to radiology reports: A multimodal machine learning approach", 2019 Digital Image Computing: Techniques and Applications (DICTA), Perth, WA, Australia, pp. 1-8, Dec. 2019.
[https://doi.org/10.1109/DICTA47822.2019.8945819]

-
V. Tiwari, M. T. Beg, and S. Goel, "Automatic generation of chest X-ray medical imaging reports using LSTM-CNN", Proc. of the International Conference on Data Science, Machine Learning and Artificial Intelligence, Windhoek Namibia, pp. 80-85, Aug. 2021.
[https://doi.org/10.1145/3484824.3484918]

-
R. Pascanu, T. Mikolov, and Y. Bengio, "On the difficulty of training recurrent neural networks", International Conference on Machine Learning, PMLR, Atlanta, Georgia, USA, pp. 1310-1318, Jun. 2013.
[https://doi.org/10.48550/arXiv.1211.5063]

-
Y. Xiong, B. Du, and P. Yan, "Reinforced transformer for medical image captioning", International Workshop on Machine Learning in Medical Imaging, Springer, Vol. 11861, pp. 673-680, Oct. 2019.
[https://doi.org/10.1007/978-3-030-32692-0_77]

-
Z. Chen, Y. Shen, Y. Song, and X. Wan, "Cross-modal memory networks for radiology report generation", arXiv preprint arXiv:2204.13258, , Apr. 2022.
[https://doi.org/10.48550/arXiv.2204.13258]

-
Z. Chen, Y. Song, T.-H. Chang, and X. Wan, "Generating radiology reports via memory-driven transformer", Proc. of the 2020 Conference on Empirical Methods in Natural Language Processing (EMNLP), Association for Computational Linguistics, pp. 1439-1449, Nov. 2020.
[https://doi.org/10.18653/v1/2020.emnlp-main.112]

-
D. You, F. Liu, S. Ge, X. Xie, J. Zhang, and X. Wu, "AlignTransformer: Hierarchical alignment of visual regions and disease tags for medical report generation", Proc. of Medical Image Computing and Computer-Assisted Intervention (MICCAI), Strasbourg, France, Vol. 24, pp. 72-82, Sep. 2021.
[https://doi.org/10.1007/978-3-030-87234-2_7]

-
Y. Zhang, H. Jiang, Y. Miura, C. D. Manning, and C. P. Langlotz, "Contrastive learning of medical visual representations from paired images and text", Machine Learning for Healthcare Conference, PMLR, Durham, NC, USA, pp. 196-219, Aug. 2022.
[https://doi.org/10.48550/arXiv.2010.00747]

-
J. Shentu and N. Al Moubayed, "CXR-IRGen: An integrated vision and language model for the generation of clinically accurate chest X-ray image-report pairs", Proc. of the IEEE/CVF Winter Conference on Applications of Computer Vision, Waikoloa, Hawaii, pp. 5212-5221, Jan. 2024.
[https://doi.org/10.1109/WACV57701.2024.00513]

-
A. E. W. Johnson, T. J. Pollard, S. J. Berkowitz, N. R. Greenbaum, M. P. Lungren, C. Deng, R. G. Mark, and S. Horng, "MIMIC-CXR, a de-identified publicly available database of chest radiographs with free-text reports", Scientific Data, Vol. 6, No. 1, pp. 317, Dec. 2019.
[https://doi.org/10.1038/s41597-019-0322-0]

-
M. Raghu, C. Zhang, J. Kleinberg, and S. Bengio, "Transfusion: Understanding transfer learning for medical imaging", Advances in Neural Information Processing Systems (NeurIPS), pp. 3347-3357, Dec. 2019.
[https://doi.org/10.48550/arXiv.1902.07208]

-
O. Vinyals, A. Toshev, S. Bengio, and D. Erhan, "Show and tell: A neural image caption generator", IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, pp. 3156-3164, Jun. 2015.
[https://doi.org/10.1109/CVPR.2015.7298935]

-
S. J. Rennie, E. Marcheret, Y. Mroueh, J. Ross, and V. Goel, "Self-critical sequence training for image captioning", IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, pp. 7008-7024, Jul. 2017.
[https://doi.org/10.1109/CVPR.2017.131]

-
J. Lu, C. Xiong, D. Parikh, and R. Socher, "Knowing when to look: Adaptive attention via a visual sentinel for image captioning", IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, pp. 375-383, Jul. 2017.
[https://doi.org/10.1109/CVPR.2017.47]

-
M. Cornia, M. Stefanini, L. Baraldi, and R. Cucchiara, "Meshed-memory transformer for image captioning", IEEE/CVF Conference on Computer Vision and Pattern Recognition, Seattle, WA, USA, pp. 10578-10587, Jun. 2020.
[https://doi.org/10.1109/CVPR42600.2020.01059]

- S. Wang, Z. Lu, X. Wang, Y. Ding, M. Li, and X. Long, "Prior knowledge enhances radiology report generation", in Proc. AMIA Summits on Translational Science, pp. 486-495, May 2022.

2019. 10 : BS degree, Dept. of Computer Science, Bacha Khan University
2023. 3 : MS degree, Dept. of Computer Science, COMSATS University
2023. 3 ~ present : Phd Student, Dept. of Intelligent Electronic and Computer Engineering, Chonnam National University
Research Interests: Machine Learning, Deep Learning, Computer Vision, Medical Imaging

2019. 8 : BS degree, Dept. of Earth and Environmental Sciences, Chonnam National University
2019. 9 ~ present: MS/Phd Student, Dept. of Intelligent Electronic and Computer Engineering, Chonnam National University
Research Interests: Machine Learning, Deep Learning, Computer Vision

1986. 2 : BS degree, Dept. of Electronic Engineering, Seoul National University
1988. 2 : MS degree, Dept. of Electronic Engineering, Seoul National University
1995. 3 ~ present: Professor, Dept. of Intelligent Electronic and Computer Engineering, Chonnam National University
Research Interests: Digital Signal Processing, Computer Vision, Machine Learning, Deep Learning